Tobacco conglomerate Philip Morris International (PMI), owner of the world’s leading cigarette brand Marlboro, has an aim that may seem strange: they are designing a smoke-free future through the development of Reduced Risk Products (RRPs).
As part of the process of developing less harmful products, PMI’s methodologies and findings are shared with scientific experts and regulators, and PMI also crowdsources independent verification of their scientific results through presenting a series of challenges on the sbv IMPROVER platform (Systems Biology Verification).
 We caught up with Stéphanie Boue, a scientist at PMI who is speaking at our CSW Arctic // Europe 2018 conference in Sweden’s Lapland on March 20-24. In her session, Stephanie will:
We caught up with Stéphanie Boue, a scientist at PMI who is speaking at our CSW Arctic // Europe 2018 conference in Sweden’s Lapland on March 20-24. In her session, Stephanie will:
- discuss concepts relevant to sbv IMPROVER and give a short history of the use of crowdsourcing in biological sciences
- cover how PMI has used crowdsourcing successfully in the past to solve a number of scientific questions in systems biology and systems toxicology
- present key learnings PMI had using crowdsourcing in science, including how to best reach out to the crowd and incentivize participation.
Stéphanie, can you please tell us about your background and how in your role at PMI you work with sbv IMPROVER?
I have a background in biotechnology (Engineering degree in Biotechnology from the Ecole Supérieure de Biotechnologie de Strasbourg (ESBS)), molecular biology, and bioinformatics, and I obtained my Ph.D. in Bioinformatics from the European Molecular Biology Laboratory (EMBL) in Heidelberg, Germany.
After a postdoc on pluripotent cells at the Center for Regenerative Medicine in Barcelona (CMRB), I joined Philip Morris International R&D as a computational biologist in 2010, just as the sbv IMPROVER concepts were put in place. I participated in the design of the very first challenge, and since then, I’ve been working on different ways to harvest the wisdom of the crowd. I now manage scientific transparency and verification aspects, including the sbv IMPROVER project, in which we organize computational challenges as well as datathons.
Did PMI use the sbv IMPROVER in the development of Reduced Risk Products (RRPs)?
sbv IMPROVER is not used directly in the development of Reduced Risk Products (RRP), but rather aims at ensuring that the methods that are used in the systems toxicology assessment of those products are state-of-the-art and reliable.
The only challenge that used data from our assessment of RRPs directly is the Systems Toxicology challenge, which aimed at verifying an exposure signature from blood transcriptomics data able to distinguish smokers from never and former smokers. It allowed us to also verify that animals and subjects who switched to a candidate RRP had profiles similar to those of animals and subjects who ceased exposure to cigarette smoke.
I see a number of challenges on sbv IMPROVER, can you walk us briefly through the Network Verification challenge and the Microbiomics challenge? What is it that you’re trying to achieve and who are the typical participants?
In our two currently open challenges, the Network Verification challenge and the Microbiomics challenge, we have different aims and target audiences.
The current iteration of the Network Verification challenge aims at refining biological network models representing the three phases of liver xenobiotic metabolism. Xenobiotic 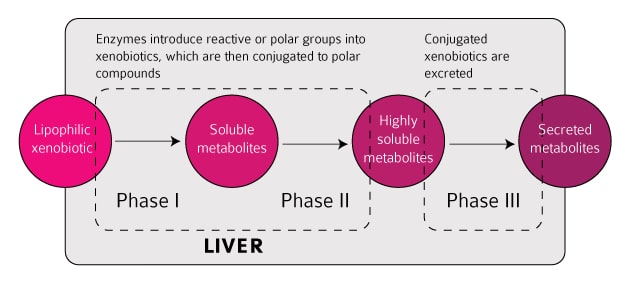 metabolism is activated in response to exposure to chemical substances foreign to the body (i.e., xenobiotics, such as toxic compounds, drugs, etc.) and is therefore relevant for multiple applications in toxicology and pharmacology. Those networks were built based on scientific literature, with one or more excerpts from a scientific publication supporting each edge in the network. The goal of the challenge is to verify the evidence supporting the network as well as to add/remove nodes and edges, as supported by the current scientific literature. Any scientist able to read and understand a scientific publication should be able to take part in the Network Verification challenge. Participation is encouraged by a leaderboard highlighting the best performers on the platform as well as by some monetary and scientific incentives.
metabolism is activated in response to exposure to chemical substances foreign to the body (i.e., xenobiotics, such as toxic compounds, drugs, etc.) and is therefore relevant for multiple applications in toxicology and pharmacology. Those networks were built based on scientific literature, with one or more excerpts from a scientific publication supporting each edge in the network. The goal of the challenge is to verify the evidence supporting the network as well as to add/remove nodes and edges, as supported by the current scientific literature. Any scientist able to read and understand a scientific publication should be able to take part in the Network Verification challenge. Participation is encouraged by a leaderboard highlighting the best performers on the platform as well as by some monetary and scientific incentives.
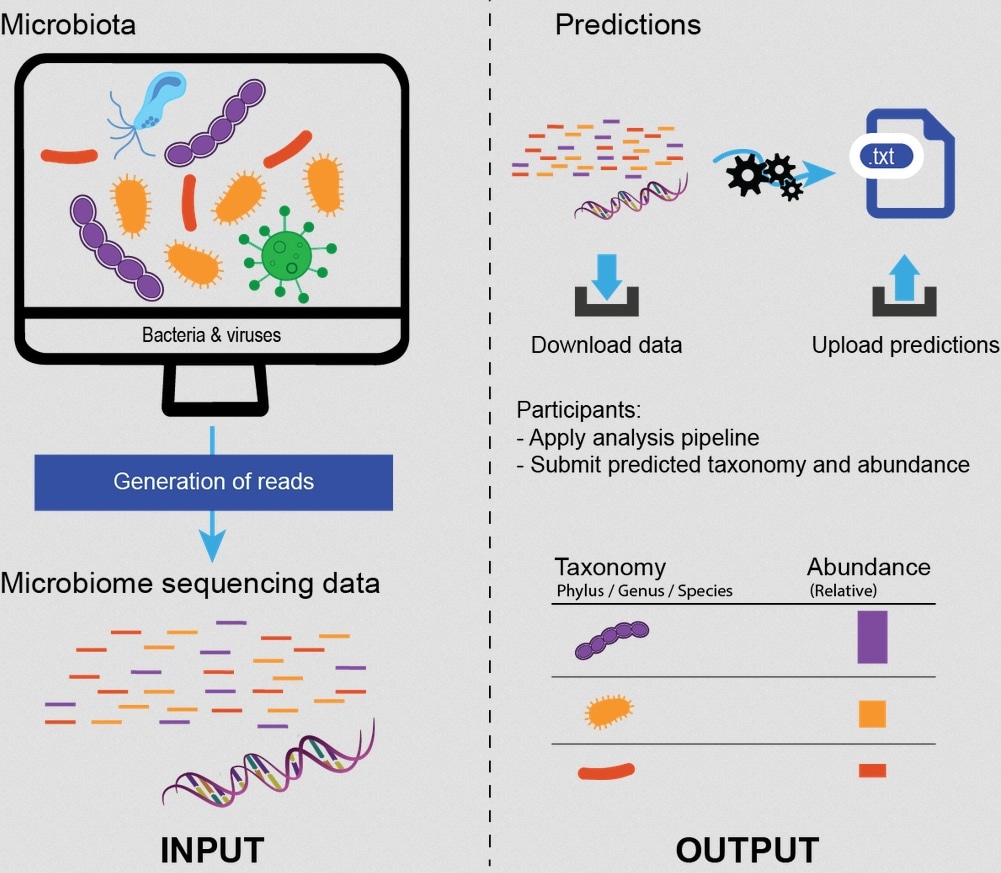
The other challenge that is currently open for participation is the Microbiomics challenge. Observing in the last few years how the microbiome is shown to play more and more significant roles in health and disease, we thought to create a challenge that would investigate computational methods used after a sample containing microbes has been sequenced. For us, this is a key first step in the investigation of the role of the microbiome in specific settings, such as certain diseases.
Hence, the Microbiomics challenge is asking computational biologists and data scientists used to analyze sequencing data to predict the bacterial composition of samples based on sequencing reads. In this challenge, participants’ predictions will be scored without bias against a gold standard. The aim is to benchmark different methods and understand which method performs best for this specific task.
Besides these two challenges, are you able to tell us what others are you looking at addressing in the future?
We plan to keep the Network Verification challenge ongoing with additional networks. We also keep monitoring scientific areas of interest and where there is a need for computational methods to be benchmarked, we will develop further challenges accordingly.
Thank you Stéphanie, we look forward to seeing you next month in Sweden.
A limited number of tickets remain available for CSW Arctic // Europe 2018 taking place inside Sweden’s Arctic Circle from 20-24 March, please check out the full agenda and we hope to see you there.





0 Comments